| 1U3C | |
| PDB ID | 1U3C |
| Japanese name | シロイヌナズナ |
| English name | Arabidopsis |
| Scientific name | Arabidopsis thaliana (L.) Heynh. |
| Protein name | Cryptochrome 1 Apoprotein |
| Gene name | Name=Cry1; Synonyms=Hy4; Orderedlocusnames=At4g08920; Gn Orfnames=T3h13.5, T3h13.14; |
| Source/Organism | Arabidopsis Thaliana |
| Compound | Molecule: Cryptochrome 1 Apoprotein; Chain: A; Fragment: Phr Domain, Residues 1-509; Synonym: Blue Light Photoreceptor; Engineered: Yes |
| Classification | Signaling Protein |
| EC number | - |
| Authors | C.A.Brautigam, B.S.Smith, Z.Ma, M.Palnitkar, D.R.Tomchick, M.Machius, J.Deisenhofer |
| Reference | C.A.Brautigam, B.S.Smith, Z.Ma, M.Palnitkar, D.R.Tomchick, M.Machius, J.Deisenhofer Structure Of The Photolyase-Like Domain Of Cryptochrome 1 From Arabidopsis Thaliana. Proc.Nat.Acad.Sci.Usa V. 101 12142 2004 |
| Key word | Photolyase, Amppnp |
| Cross reference | SWS:CRY1_ARATH,Q43125 EMBL; S66907; AAB28724.1; -. EMBL; S66909; AAB28725.2; ALT_FRAME. EMBL; AF128396; AAD17364.1; ALT_SEQ. EMBL; AL161513; CAB78016.1; ALT_SEQ. EMBL; AF361588; AAK32756.1; -. PIR; S39058; S39058. HSSP; P00914; 1DNP. IntAct; Q43125; -. InterPro; IPR002081; DNA_photolyase_1. InterPro; IPR006050; DNA_photolyase_N. InterPro; IPR005101; FAD_binding_7. InterPro; IPR006051; FAD_binding_N. Pfam; PF00875; DNA_photolyase; 1. Pfam; PF03441; FAD_binding_7; 1. PRINTS; PR00147; DNAPHOTLYASE. ProDom; PD004390; FAD_binding_N; 1. PROSITE; PS00394; DNA_PHOTOLYASES_1_1; 1. PROSITE; PS00691; DNA_PHOTOLYASES_1_2; 1. |
| Experimental method | X-Ray Diffraction, Resolution: 2.60 Angstrom, R-Factor: 0.205, R-Free: 0.254 |
| Sequence characteristics | Length: 681 AA, Molecular weight: 76793 Da |
| Sequence | MSGSVSGCGSGGCSIVWFRRDLRVEDNPALAAAVRAGPVIALFVWAPEEEGHYHPGRVSR WWLKNSLAQLDSSLRSLGTCLITKRSTDSVASLLDVVKSTGASQIFFNHLYDPLSLVRDH RAKDVLTAQGIAVRSFNADLLYEPWEVTDELGRPFSMFAAFWERCLSMPYDPESPLLPPK KIISGDVSKCVADPLVFEDDSEKGSNALLARAWSPGWSNGDKALTTFINGPLLEYSKNRR KADSATTSFLSPHLHFGEVSVRKVFHLVRIKQVAWANEGNEAGEESVNLFLKSIGLREYS RYISFNHPYSHERPLLGHLKFFPWAVDENYFKAWRQGRTGYPLVDAGMRELWATGWLHDR IRVVVSSFFVKVLQLPWRWGMKYFWDTLLDADLESDALGWQYITGTLPDSREFDRIDNPQ FEGYKFDPNGEYVRRWLPELSRLPTDWIHHPWNAPESVLQAAGIELGSNYPLPIVGLDEA KARLHEALSQMWQLEAASRAAIENGSEEGLGDSAEVEEAPIEFPRDITMEETEPTRLNPN RRYEDQMVPSITSSLIRPEEDEESSLNLRNSVGDSRAEVPRNMVNTNQAQQRRAEPASNQ VTAMIPEFNIRIVAESTEDSTAESSSSGRRERSGGIVPEWSPGYSEQFPSEENRIGGGST TSSYLQNHHEILNWRRLSQTG |
| Image | 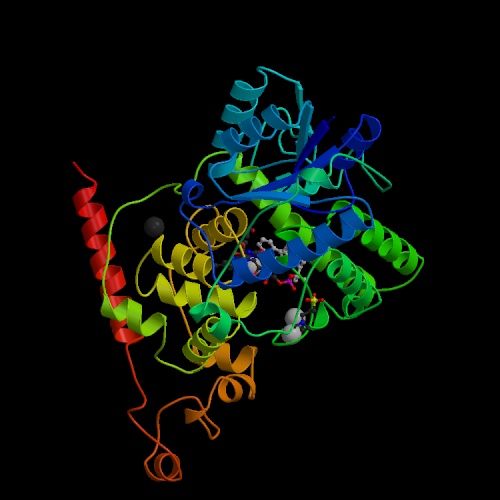 |