| FEA2 | |
| Project ID | 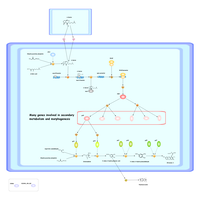 FEA2 [Protein] |
| Project Theme | Structure-based functional analysis of key enzymes that can be applied to production of antibiotics and other useful compounds |
| Project Theme (short) | Enzymes for useful compounds |
| Principal Investigator | Yasuo Ohnishi |
| Affiliation | Graduate School of Agricultural and Life Sciences, The University of Tokyo |
| Backgrounds | - Actinobacteria are well known as secondary metabolite producers including antibiotics - Elucidation of the biosynthesis pathway of the secondary metabolites could lead to efficient productions of useful materials |
| Highlights | - Structures and functions of a variety of proteins involved in the biosynthesis pathway of the secondary metabolites of actinobacteria have been revealed - We have determined structures of several enzymes including synthase for curcumin, a medicinal ingredient of turmeric - C-nitrosation pathways in nitrosobenzamide biosynthesis of actinobacteria have been deciphered |
| Outline | While primary metabolites such as amino acids, nucleic acids, and lipids are essential for biological activities, secondary metabolites are organic compounds that are not absolutely required for the survival of the organism. Examples of the products include antibiotics and pigments. Actinobacteria are well known as secondary metabolite producers including antibiotics and hence of high pharmacological and commercial interest. In this project, we will investigate enzymes involved in the biosynthesis pathway of one of the secondary metabolites, grixazone, in Streptomyces griseus. Specifically, we will carry out functional and structural characterization of enzymes GriI, GriH and GriF which can be utilized for the production of useful materials. Since the grixazone biosynthesis is under the control of a hormone A-factor, A-factor biosynthesis enzyme (AfsA) and a transcription factor AdpA will also be analyzed. Finally, curcumin synthase (CUS) discovered in rice is our research target in terms of biological functions of Turmeric (Curcuma longa). |
| Review | J. Bacteriol. Curr Opin Chem Biol |
| CSML File | FEA2.csml |