| FBB4 | |
| Project ID | 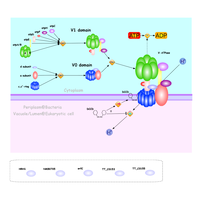 FBB4 [Protein] |
| Project Theme | Towards structure-based design of novel inhibitors for V -ATPase |
| Project Theme (short) | V-ATPase inhibitor design |
| Principal Investigator | So Iwata |
| Affiliation | Graduate School of Medicine, Kyoto University |
| Backgrounds | - Membrane transport proteins exist and span the membrane across which they transport ions and molecules - V-ATPases are highly conserved enzyme and acidify a wide array of intracellular organelles and pump protons across the membranes - V-ATPases have been implicated in osteoporosis and tumor cell invasion |
| Highlights | - Structures and functions of the rotor ring (K-ring) of V-ATPase and its complex with an inhibitor (DCCD) have been revealed - We determined the crystal structure of the DF complex of V-ATPase - The structure of a bile acid transporter ASBT, a target for hypercholesterolaemia drugs, was solved - We reported the crystal structure of hydantoin membrane transporter of an inward-facing conformation |
| Outline | The pH of different cellular compartments is tightly regulated. V-ATPases are highly conserved enzyme and acidify a wide array of intracellular organelles and pump protons across the plasma membranes. V-ATPases couple the energy of ATP hydrolysis to proton transport across membranes of eukaryotic cells. We found eukaryotic V-ATPase-like proteins in some prokaryotic cell membranes. V-ATPases consists of the Vo domain (membrane associated subunits) and the V1 domain (peripherally associated subunits). The V1 domain is responsible for ATP hydrolysis, whereas the Vo domain is for proton translocation. Plasma membrane V-ATPases have been implicated in osteoporosis and tumor cell invasion, emerging as potential targets in the treatments of osteoporosis and cancer. In this project, we try to identify compounds which inhibit the activity of human V-ATPases by utilizing prokaryotic V-ATPases as model systems. |
| Review | Nat Commun |
| CSML File | FBB4.csml |