| FBA2 | |
| Project ID | 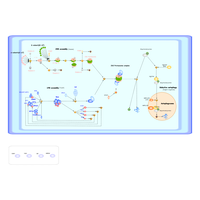 FBA2 [Protein] |
| Project Theme | Structural basis for dynamic formation and mechanistic actions of huge and complicated proteolytic machinery |
| Project Theme (short) | Proteolytic system: proteasome |
| Principal Investigator | Keiji Tanaka |
| Affiliation | The Tokyo Metropolitan Institute of Medical Science, Tokyo Metropolitan Organization for Medical Research |
| Backgrounds | - Cells possess the protein degrading machinery called proteasome - The proteasome is a huge barrel-like protein complex and its assembly process and structure remain unclear - Aberrant proteasome activity has been implicated in several disease such as Alzheimer's disease and cancer |
| Highlights | - Structures and functions of several proteasome-dedicated chaperones involved in the assembly of the 20S proteasome have been revealed - Efforts are in progress to elucidate the entire 26S proteasome structure |
| Outline | Proteasome (26S Proteasome) degrades unneeded or damaged proteins by proteolysis to regulate the concentration of particular proteins in cells. The proteasomal degradation pathway is essential for many cellular processes, including the regulation of metabolism and immune response. Decreased proteasome activity has been suggested as a cause of several neural diseases such as Parkinson's disease and Alzheimer's disease. The 26S proteasome, a huge protein complex comprising of about 100 molecules, contains one 20S core particle structure and two 19S regulatory caps. The core is hollow and provides an enclosed cavity in which proteins are degraded; openings at the two ends of the core allow the target protein to enter. Since we already succeeded in the structural analysis of the central catalytic part (20S core particle), our next target is 19S regulatory particle. Several proteasome-dedicated chaperones are involved in the efficient and correct assembly of the 20S proteasome. These chaperones help the initiation and progression of the assembly process by transiently associating with proteasome precursors. |
| Review | Nat. Rev. Mol. Cell Biol. J. Biochem. |
| CSML File | FBA2.csml |